Data source integration
| Author(s) |
|
| Reviewers |
|
OverviewQuestions:
Objectives:
How can I write a tool that can import data into Galaxy from an external database?
What are “data sources” and how do they function?
Is there any ready-to-use example?
Time estimation: 10 minutesSupporting Materials:Published: Sep 30, 2016Last modification: Oct 18, 2022License: Tutorial Content is licensed under Creative Commons Attribution 4.0 International License. The GTN Framework is licensed under MITpurl PURL: https://gxy.io/GTN:T00114version Revision: 15
Data Source Integration
An important goal of Galaxy is scalability. A major bottleneck when it comes to analysis of big data sets is the time and space it takes of copying these data sets.
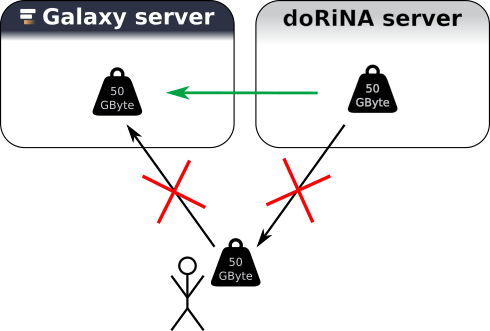
Galaxy provides an interface such that it can communicate with other servers to get data directly into the Galaxy environment of a user without the need of “downloading” the data. In this hands on, we will use the resource from DoRiNA Server (Blin et al. 2014). but the main point about this short section is: if you have a data source which you think is very important for your research with Galaxy let us know!
Hands On: Hands on!
- Create a new history called “doRiNA”
- Go to Get Data::doRiNA search
- Choose hg19 from the drop-down list -> Search Database
- Leave everything as is and choose from the Regulators (set A) drop-down list “hsa-let-7astar-CLASH” -> Search doRiNA
- Use the “Send to Galaxy” button
- Notice the new History Item
You can see the tutorial section of the DoRiNA website for more detailed examples. That was very easy for all of you! If you want your database of choice to be accessible as easy as this let us know!